- This event has passed.
Second Philosophy of Cancer Biology Workshop, with Jerome Galon and Joel Brown (Pey Berland, Bordeaux, France)
20 January 2020 - 21 January 2020

January 20th-21st, 2020
University of Bordeaux (France)
Campus Pey Berland (35 Place Pey Berland 33000 Bordeaux)
The University of Bordeaux, the CNRS, ImmunoConcEpT, and PhilInBioMed host the second international workshop on the Philosophy of Cancer Biology in Bordeaux, France, organized by Wiebke Bretting, Sara Green, Lucie Laplane, Maël Lemoine, and Thomas Pradeu. The workshop is funded by Thomas Pradeu’s ERC-funded project IDEM.

Context
Cancer is one of the main causes of death globally according to the World Health Organization. The biological complexity and heterogeneity of this disease (or group of diseases) make it very difficult to apprehend, control, and cure. For a long time, cancer has been little studied by philosophers of science. Most of the work in the humanities and the social sciences has focused on the social, anthropological, psychological, and ethical dimensions of cancer. Yet cancer is now becoming increasingly an object of study for philosophers of biology and philosophers of medicine. In particular, the scientific explanation, definition, classification and prediction of cancer as a biological and medical phenomenon face many epistemological challenges. Cancer research raises a host of experimental, theoretical, and conceptual issues that connect with most, if not all, the domains of today’s biology and medicine.
The main goal of this workshop is to provide a forum where philosophers of biology and philosophers of medicine meet to discuss the biological and medical science of cancer.
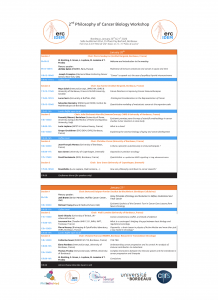
Program and list of speakers
Program (please click to enlarge)
Plenary speakers
Jerome Galon (Research Director at INSERM, Head of the Laboratory of Integrative Cancer Immunology, Paris, France) – Multiverse of immune contexture and cancers in space and time
Joel Brown (Senior Member, Moffitt Cancer Center, USA) – Using Principles of Ecology and Evolution to Define, Understand and Treat Cancer
Invited speakers
Sebastien Benzekry (INRIA team MONC, Institut de Mathématiques de Bordeaux) – Quantitative modeling of metastasis: cancer at the organism scale
Sara Green (Section for History and Philosophy of Science, Department of Science Education, University of Copenhagen, Denmark) – Organoids in precision oncology
Lucie Laplane (IHPST, CNRS & University Paris 1; Institut Gustave Roussy, France) – What is a clone?
Jean-François Moreau (University of Bordeaux, France) – Is there a place for autoimmunity in immunotherapies ?
Pierre Nassoy (Bioimaging & OptoFluidics laboratory, CNRS, Bordeaux, France) – Organoids: a brain teaser in physics of Active Matter and more than just a toy-model in Oncology?
Samir Okasha (University of Bristol, UK) (via video-conference) – Cancer, evolutionary conflict, and levels of selection
Thomas Pradeu (ImmunoConcept, UMR5164, CNRS & University of Bordeaux; IHPST, CNRS & University Paris 1, France) – Complex interactions between the immune system and the microbiota in cancer progression and therapies
Maya Saleh (ImmunoConcept, UMR5164, CNRS & University of Bordeaux, France & McGill University, Canada) – Future Directions in Improving Cancer Immunotherapies
David Santamaria (PI “Novel mediators in lung oncogenesis”, IECB, Bordeaux, France) – Quantitative vs qualitative MAPK signalling in lung adenocarcinoma
Catherine Sawai (INSERM U1218, Bordeaux, France) – In vivo models of normal and leukemic hematopoiesis
Selected speakers
Joseph Crompton (Memorial Sloan Kettering Cancer Center, New York, USA) – “Cancer” as speech act: the case of papillary thyroid microcarcinoma
Gregor Greslehner (ERC IDEM, CNRS, Bordeaux, France) – Explaining the common biology of aging and cancer development
Laurence Huc (INRA, Toulouse, France) – What is a carcinogen? bridging the gap between basic biology and regulatory toxicology
Andrea Pensotti (University of Rome, University Campus Bio-Medico of Rome, Italy), Marriano Bizzarri (Sapienza University, Italy), and Marta Bertolaso (University of Rome, University Campus Bio-Medico of Rome, Italy) – Epistemic premises pave the way of scientific methodology in cancer studies: tumor reversion as case study
Elena Rondeau (University of Bordeaux, France) – Understanding cancer progression and its control: An analysis of immune contribution to metastasis
Lucas Serra (University at Buffalo, USA) – Ontological Considerations in the Representation of Cancer
Michael J. Young (Harvard Medical School, USA) – Epistemic Upshots of the Genomic Turn in Cancer Care: Lessons from Neuro-Oncology
Abstracts
Plenary speakers
Jerome Galon (Research Director at INSERM, Head of the Laboratory of Integrative Cancer Immunology, Paris, France)
Multiverse of immune contexture and cancers in space and time
To-date, the evaluation of the prognosis of cancer patients relied on the anatomic extent of tumor (TNM-classification). However, this classification provides limited prognostic information and does not predict response to therapy. We redefined cancer by integrating the immune system to transfer cutting-edge medicine to the patients. We have previously shown that tumors from human colorectal cancer with a high-density of infiltrating memory and effector-memory T-cells are less likely to disseminate to lympho-vascular and perineural structures and to regional lymph-nodes. We also demonstrated the critical tumor-microenvironment parameters determining the dissemination to distant metastasis. We found that the combination of immune parameters associating the nature, the density, the functional immune orientation and the location of immune cells within the tumor was essential to accurately define the impact of the local host-immune reaction on patients’ prognosis. We defined these parameters as the “immune contexture”. We characterized the immune landscape within human tumors, and showed the importance of several adaptive immune cells. We described the immunophenotype and antigenome associated with immune escape mechanisms and demonstrated mechanisms associated with pre-existing and proliferating intratumoral T-cells. Based on the immune contexture, a standardized, simple and powerful digital-pathology-based immune stratification-system, termed “Immunoscore”, was delineated having a prognostic power superior to that of the currently used cancer staging-system. Tumor invasion parameters were statistically dependent on the host-immune reaction. A worldwide consortium validated the prognostic value of the consensus Immunoscore, using a standardized-assay. We have demonstrated the significant role of Immunoscore and immunoediting in affecting metastatic dissemination in space and time. We hence proposed a “parallel immune selection model” of tumor evolution incorporating the effects of the immune system in shaping and driving metastatic spread. Further analyses revealed a large inter- and intra-metastatic immune heterogeneity. The same study revealed how Immunoscore and T and B cell score (TB score) from the least-infiltrated metastasis are the most associated with survival. Thus, tumor progression, invasion and recurrence are dependent on pre-existing immunity and on Immunoscore. Finally, novel concepts underpinning tumor evolution at the pre-cancerous stages will be advocated.
Joel Brown (Senior Member, Moffitt Cancer Center, USA)
Using Principles of Ecology and Evolution to Define, Understand and Treat Cancer
“You have cancer.” What unfortunate words. To patient, family and friends, cancer brings a maelstrom of emotions including fear and hope. It can be a horrific disease of genetic mutations and unregulated proliferation. But, cancer is much more, and knowing this can empower the patient and suggest new therapies. Definitions of cancer that focus on genes, proliferation or the capacity to invade other tissues may describe what cancer does but they do not describe what cancer is. Cancer may best be defined as a speciation event. A cell lineage goes from being part of the whole body program to becoming its own program – it becomes the unit of natural selection. Thus, cancer cells inhabit a tumor ecosystem where they experience much the same hazards and opportunities present in the ecology of any creature. Furthermore, like nature, they evolve adaptations to better acquire resources, avoid the hazards of the immune system, and occupy new spaces and organs of the patient. We know that the failure of therapy lies in the capacity of cancer cells to evolve resistance. Yet, despite vast sums of money, legions of researchers, and millions of publications, about 1 in 4 of those living in the EU and USA can, at present, expect to die from cancer. Survival rates from late stage, metastatic cancers have changed little in years. Why? Perhaps cancer is uniquely complex, uniquely intractable compared to other ailments, or simply requires more money, brainpower and research. Here, I would suggest that our failure to conceptualize cancer properly has stymied progress. An eco-evolutionary perspective offers new insights, and suggests novel therapeutic strategies. Such therapies aim to use drugs more sparingly and judiciously. We can and should anticipate and steer the cancer cells’ evolution. In this way, otherwise incurable cancers may be managed as a livable, chronic disease, or better yet, cured by beating cancer at its own ecological and evolutionary “chess” game.
Invited speakers
 Sebastien Benzekry (INRIA team MONC, Institut de Mathématiques de Bordeaux)
Sebastien Benzekry (INRIA team MONC, Institut de Mathématiques de Bordeaux)
Quantitative modeling of metastasis: cancer at the organism scale
In the majority of solid cancers, secondary tumors (metastases) are the main cause of death. Determining the burden of invisible metastases at diagnosis and predicting how they would respond to treatments is a crucial challenge in the clinic, for multiple cancer types. Indeed, this would allow personalization of the extent of therapeutic interventions, for instance in the perioperative setting. I will present research efforts towards the establishment of such a predictive computational tool of metastatic development, with emphasis on the quantitative calibration of models to empirical data (experimental and clinical). The general framework is based on a physiologically-structured partial differential equation for the time dynamics of a population of metastases. Results will be presented in two clinical settings: brain metastasis from non-small cell lung cancer and early-stage breast cancer. In the first application, comparison of different models – based on different biological hypotheses about dissemination and growth – indicates periods of dormancy of the order of several months. In the second application, a combination of machine learning techniques and advanced statistical learning methods allow individualized predictions of the model’s parameters from data available at diagnosis. In turn, this allows patient-specific prediction of the time to metastatic relapse. Together, these results represent a step towards the integration of mathematical modeling as a predictive tool for personalized oncology.
References
Benzekry, S., Tracz, A., Mastri, M., Corbelli, R., Barbolosi, D., & Ebos, J. M. L. (2016). Modeling Spontaneous Metastasis following Surgery: An In Vivo-In Silico Approach. Cancer Research, 76(3), 535–547. http://doi.org/10.1158/0008-5472.CAN-15-1389
Bilous, M., Serdjebi, C., Boyer, A., Tomasini, P., Pouypoudat, C., Barbolosi, D., et al. (2019). Quantitative mathematical modeling of clinical brain metastasis dynamics in non-small cell lung cancer. Scientific Reports, 9(1), 13018.
 Sara Green (Section for History and Philosophy of Science, Department of Science Education, University of Copenhagen, Denmark),
Sara Green (Section for History and Philosophy of Science, Department of Science Education, University of Copenhagen, Denmark),
Organoids in precision oncology
A common problem in oncology is that cancer tumors growing in the body of individual patients often respond differently to medical treatments, compared to standardized cancer cell lines or mouse models. Drugs that perform well in research on cancer models often fail in clinical trials and are hence difficult to translate to clinical practice (Drost & Clevers 2018). To account for the problem of tumor heterogeneity, organoids based on tumors from cancer patients are currently being developed for a variety of purposes. Tumor organoids are three-dimensional tissue cultures developed from stem or cancer cells. Growing 3D cultures or organ-fragments is not new in cancer biology, but tissue culture techniques can now be combined with technologies for advanced molecular profiling and imaging analysis (Shen 2018). Many have described organoids as revolutionary for the study of cancer and other diseases (Akkerman& Defize 2017), and Nature recently endowed organoids as the “Method of the Year” (Nature Editorial 2018). Organoids are becoming more widespread in cancer research and drug testing, and patient-derived organoids are also explored for their capacity as pre-clinical models to enable patient-specific drug screening (Vlachogiannis et al. 2018). This paper explores how the use of patient-derived organoids impacts research and diagnostic practice. We combine philosophical and ethnographic analysis to compare how translational potentials and challenges of organoids are assessed in scientific discussions and experienced in clinical practices.
Organoids have characteristics that both resemble and differ from more traditional models in cancer research, such as 2D cancer cell cultures and mouse models. Unlike 2D cellular models, tumor organoids grown in a 3D matrix recapitulate some features of the tumor micro-environment and retain biological characteristics of tumors better than cancer cell lines (Xu et al. 2018). Moreover, in their capacity as patient-derived (rather than standardized) models, organoids account for heterogenous features that may allow for better understanding and prediction of cancer development and drug response in specific patients or patient groups. Parallel studies of drug response in organoids grown from tumors and normal tissue can help establish which drugs primarily target cancer cells without harming normal cells. In addition, organoids allow for the study of the evolvability of finer-grained cancer types. We present examples of early results from such studies and reflect on the associated translational potentials and challenges. Our analysis shows that although organoids and other patient-derived models are sometimes framed as a technique that allows for direct inferences of drug response, their evidential status remains a controversial issue in the field. At present, tumor organoids can be interpreted as epistemic objects, i.e., as unstable research entities that embody aspects we do not yet know and therefore drive science forward in their function as “question generating machines” (Rheinberger 1997). We show that in the context of tumor organoids, open-ended questions about their evidential status are intimately connected to uncertainties about the nature of cancer itself.
One important consideration is whether organoids adequately capture features typically studied in mouse models. Compared to (patient-derived) murine models, organoids are easier and less resource-demanding to develop, and can be seen as simpler models that allow for high-throughput drug screening (Yang et al. 2018). At the same time, however, organoids lack important elements of the natural environment of cancer tumors, such as blood vessels, immune cells, and other stromal components that are known to influence tumors beyond the tumor microenvironment (Lapane et al. 2019). A specific concern is therefore the extent to which organoids grown in vitro retain the characteristics of in situ tumors, particularly whether growth conditions impact the histology, genetic profile, and drug response of tumor organoids.
Another consideration is how precision oncology balances the persistent tension between standardization and variation in medicine. The ability of organoids to account for heterogeneity (between patients and between patient tumors at different spatial and temporal scales) is typically seen as a positive feature. However, in a context where each tumor is considered as unique and internally heterogenous, there may be trade-offs between the variability of the experimental system and the reproducibility of experimental results (Huch et al. 2017). It is therefore unclear to what extent organoids present an alternative to traditional model organisms or standardized cells, and what level of variation research should aim for if the aim is to infer molecular action mechanisms (Lang 2019). In practice, accounting for patient-specific variation via tumor organoids also requires that other aspects are standardized (e.g., growth factors, structural constraints in the 3D matrix, and drug doses), which raises questions about what aspects of medicine are possible or most useful to “personalize“. These discussions relate to the broader debate about what evidence means in the context of hyper-stratified disease classes in precision oncology. Rather than primarily asking whether organoids adequately represent the patient tumor, we explore how practitioners in the field balance the benefits and limitations of different model systems. This involves how organoid research is related to other sources of evidence, and how questions about evidence-status are tied to social and ethical implications in translational contexts.
We suggest that organoids can further be interpreted as collaborative things, i.e., as entities that expand translational collaborations by facilitating new forms of exchanges across new contexts (Michael 2005; Davies 2012). Organoids provide an entry into infrastructures and collaborations that affect the relations between cancer research, clinical practice, and the biomedical industry. Organoids constitute infrastructures to generate living biobanks of cryopreserved organoids as a resource for research, drug development, and drug screening (Drost & Clevers 2018). Moreover, organoids are at the center of translational projects that establish new and tighter connections between research and clinical practice.
As part of an ethnographic research project, we have followed a preclinical trial, where organoids are used to guide decisions on experimental cancer treatments in a phase-I clinic based on genetic profiling. Interesting findings from this context involve practical and temporal constraints that are often absent from scientific review papers. Unlike the war rhetoric of rapidly expanding cancer cells, patient-derived organoids (especially from biopsies) are in this context often perceived as fragile entities that are difficult to grow within the clinically meaningful time window. Moreover, while organoids generate new hope for terminal cancer patients by presenting an intuitive and personalized form of evidence, the interpretation of drug response is far from straightforward. For oncologists, the possibility to attain new forms of evidence therefore also entails a confrontation with new types of uncertainties – as well as new ethical responsibilities to inform patients about these. In this context, organoids transform terminal cancer patients into what we may call precision patients, i.e., as epistemic objects on their own whose results will help answer questions about the potential of precision oncology.
References
Akkerman, N., & Defize, L. H. (2017). Dawn of the organoid era: 3D tissue and organ cultures revolutionize the study of development, disease, and regeneration. Bioessays, 39(4), 1600244.
Davies, G. (2012). What is a humanized mouse? Remaking the species and spaces of translational medicine. Body & Society, 18(3-4), 126-155.
Drost, J., & Clevers, H. (2018). Organoids in cancer research. Nature Reviews Cancer, 18(7), 407.
Huch, M., Knoblich, J. A., Lutolf, M. P., & Martinez-Arias, A. (2017). The hope and the hype of organoid research. Development, 144(6), 938-941.
Lang, S. (2019). What are the pros and cons of using organoids? Drug target review, available online: https://www.drugtargetreview.com/article/48244/what-are-the-pros-and-cons-of-using-organoids/.
Laplane, L., Duluc, D., Bikfalvi, A., Larmonier, N., & Pradeu, T. (2019). Beyond the tumour microenvironment. International journal of cancer, available online: https://doi.org/10.1002/ijc.32343
Michael, M., Wainwright, S. P., & Williams, C. (2005). Temporality and Prudence: On Stem Cells as” Phronesic Things”. Configurations, 13(3), 373-394.
Nature Editorial (2018). Method of the Year 2017: Organoids. Nature methods, 15(1), 1.
Rheinberger, H. J., & Fruton, J. S. (1997). Toward a history of epistemic things: Synthesizing proteins in the test tube (pp. 75-90). Stanford, CA: Stanford University Press.
Shen, H. (2018). Core Concept: Organoids have opened avenues into investigating numerous diseases. But how well do they mimic the real thing? Proceedings of the National Academy of Sciences, 115(14), 3507-3509.
Vlachogiannis, G., Hedayat, S., Vatsiou, A., Jamin, Y., Fernández-Mateos, J., Khan, K., … & Rata, M. (2018). Patient-derived organoids model treatment response of metastatic gastrointestinal cancers. Science,359(6378), 920-926.
Xu, H., Lyu, X., Yi, M., Zhao, W., Song, Y., & Wu, K. (2018). Organoid technology and applications in cancer research. Journal of hematology & oncology, 11(1), 116.
Yang, H., Sun, L., Liu, M., & Mao, Y. (2018). Patient-derived organoids: a promising model for personalized cancer treatment. Gastroenterology Report, 6(4), 243–245.
 Lucie Laplane (IHPST, CNRS & University Paris 1; Institut Gustave Roussy, France),
Lucie Laplane (IHPST, CNRS & University Paris 1; Institut Gustave Roussy, France),
What is a clone?
Tumours are composed of heterogeneous cells (intratumor heterogeneity, ITH). This heterogeneity makes cancer cells difficult to target, and contribute to therapy escape and relapses. The clonal evolution model depicts the dynamical process of emergence, growth, decline, or disappearance of the clones constituting a tumour through space and time. Understanding these dynamics allows to avoid some pitfalls (such as selection of resistant clones) and built innovative therapeutic strategies accounting for these dynamics (such as adaptive therapies that maintains competition between clones). However, the very notion of clone is much more ambiguous than it looks like. The definition of a clone is straightforward and refers to a group of cells sharing a common identity inherited from a common cell of origin. But no two cells are identical in every respect, not even at the genetic level. Thus, a clone is always defined with respect to a particular trait or characteristic. Mutations are the most usual traits of interest used to describe clones, but choices can vary from the study of the whole genome to a few selected genes, or alternatively other traits can be used such as epigenetic alterations. In this talk I will discuss how technical and ontological choices impact the notion of clone, and their consequence for cancer treatment.
(1) Panel sequencing versus whole exome or whole genome sequencing, and the depth of sequencing drastically change the degree of ITH, and the number of clones depicted in a given tumour. It also changes the number of healthy individuals carrying cancer associated mutations. For example, with a threshold of variant allele frequency of 0.02%, around 10% of 70-years olds healthy individual have a clone containing a mutation typical of blood cancer, raising the question of whether these people should be treated to avoid potential transformation. With a threshold 0.0003%, 95% of 50-60 years old individuals have DNMT3A and/or TET2 mutations in their blood, questioning the relevance of these mutations in cancer transformation.
(2) Clonal evolution studies largely rely on the assumption that clonal architecture is driven by the impact of mutations on cell fitness. I will discuss confounding factors and alternative explanations of clonal dynamics, in particular epigenetic alterations and stem cells dynamics.
Altogether, these issues raise a general question about who, when, and what (which clones?) should be treated.
 Jean-François Moreau (University of Bordeaux, France)
Jean-François Moreau (University of Bordeaux, France)
Is there a place for autoimmunity in immunotherapies ? (JF Moreau, M Lemoine, E Laconi, T Pradeu)
Immunotherapies seem to be the magic bullet to cure patients with some types of cancer. Whereas the mutation load is coming to mind and is sustained by many studies, to explain the increased immune response toward the tumor, it should not be overlooked that ipilimumab (targeting CD152/ CTLA-4) or nivolumab (targeting PD-1) or atezolizumab (targeting PD-L1) among others, do alter the fine tuning of the level of autoreaction within the immune system which is deeply rooted into the functionning of the adaptive immune system. Usually considered as the underpinnings of the tolerance mechanisms, we will explain the basis of it and its potential consequences in the natural evolution of cancer and in the response to the today immunotherapies in the field of cancer.
 Pierre Nassoy (Bioimaging & OptoFluidics laboratory, CNRS, Bordeaux, France)
Pierre Nassoy (Bioimaging & OptoFluidics laboratory, CNRS, Bordeaux, France)
Organoids: a brain teaser in physics of Active Matter and more than just a toy-model in Oncology?
The current challenge in cancer biology and regenerative medicine is to engineer cellular complexity into multicellular tumor spheroids and organoids in a controlled manner. We have developed a microfluidic method that allows to produce organoids in a versatile and high throughput format. Our method is especially well suited to investigate the impact of extracellular matrix-cells interactions and to decipher key mechanotransduction cues. As physicists, we have first explored the simplest and less physiological cases to unveil the unique material properties of organoids as a prototypical example of active soft matter. Then, in the context of oncology, we aim at engineering realistic tumor models amenable to quantification and elucidation of the main determinants of tumor progression. On the basis of this dual viewpoint and our experimental data, with the risk of breaking down open doors, we will highlight that, not only the spatial environment, but also the time scale at which (tumor) organoids need to be investigated is of primary importance.
 Samir Okasha (University of Bristol, UK) (via video-conference)
Samir Okasha (University of Bristol, UK) (via video-conference)
Cancer, evolutionary conflict, and levels of selection
In the evolutionary biology literature, cancer is often regarded as a form of conflict between cells and organisms. On this view, the proliferation of cancer cells represents a form of within-organism selection, in which selfish cancer cells ruthlessly seek their own short-term proliferation, to the detriment of the whole organism, who evolve mechanisms to prevent it. Thus the framework of multi-level selection theory is often applied to cancer. Cancer cells are described as “cheats”, pursuing their own interests at the expense of the collective; and an analogy is drawn between the mechanisms by which organisms fight cancer (e.g. apoptosis) and the suppression mechanisms found in social insect colonies to enforce cooperation. In two recent articles, A. Gardner (2015) and M. Shpak and J. Liu (2016) argue that cancer is not a bona fide form of multi-level selection, that cancer cells should be not regarded as “selfish” or “cheats”, and that the analogy between anti-cancer adaptations and suppression mechanisms in social groups is misleading. Their argument turns on the fact that genetic mutations in cancer cells are an evolutionary dead-end, as they are not transmitted to the next generation. I analyze this argument in detail. Though Gardner and Shpak & Liu make an important point, I consider two possible ways of salvaging the idea that cancer represents a form of evolutionary conflict between cells and organisms. The first draws on some of the Leo Buss’s ideas in The Evolution of Individuality. The second draws on the idea that cancer represents a form of atavism, or reversion to an ancestral mode of life.
 Thomas Pradeu (ImmunoConcept, UMR5164, CNRS & University of Bordeaux; IHPST, CNRS & University Paris 1, France),
Thomas Pradeu (ImmunoConcept, UMR5164, CNRS & University of Bordeaux; IHPST, CNRS & University Paris 1, France),
Complex interactions between the immune system and the microbiota in cancer progression and therapies
It is increasingly recognized that almost all organisms are hosts to myriads of microbes, which often play functional roles in the host. Recent research has revealed that the microbiota (including viruses) could have a strong impact on cancer development (Zitvogel et al., 2018). The microbiota plays a role in carcinogenesis, both locally and at distant sites. Furthermore, the microbiota can influence how the host responds to anti-cancer therapies. Several of these therapies, including chemotherapy and several immunotherapies, show reduced efficacy in germ-free mice as well as in mice treated with broad-spectrum antibiotics, or in mice lacking specific bacteria that stimulate the immune system (Iida et al., 2013; Sivan et al., 2015; Vétizou et al., 2015; Viaud et al., 2013). In parallel, it was observed in patients that certain bacteria had a positive impact on host responses to some immunotherapies. Initial correlational observations were then confirmed causally, by fecal microbiota transplantation (FMT) from patients to mice, which suggested that “responding” and “non-responding” phenotypes could be transferred from humans to mice, with some particular bacteria playing a crucial role in this process (Gopalakrishnan et al., 2018; Matson et al., 2018; Routy et al., 2018).
This research raises many interesting issues, at the interface between biology, medicine, and philosophy:
i) Should the influence of the microbiota on tumor development be considered as one illustration among others of the increasingly acknowledged role of the tumor microenvironment (TME) (Laplane et al., 2018, 2019)?
ii) Do such studies demonstrate mere correlation, or causality? (Thomas and Jobin, 2015) What are the mechanisms underlying the influence of the microbiota on tumor development, and what is the role of the immune system in this process?
iii) Is it possible to better characterize the specific bacteria that seem to have a positive role in host’s response to cancer treatment?
iv) More generally, what does the impact of the microbiota on cancer development tell us about our definition of cancer, and about emerging “ecosystemic” approaches in the clinic?
References:
Gopalakrishnan, V., Spencer, C.N., Nezi, L., Reuben, A., Andrews, M.C., Karpinets, T.V., Prieto, P.A., Vicente, D., Hoffman, K., Wei, S.C., et al. (2018). Gut microbiome modulates response to anti–PD-1 immunotherapy in melanoma patients. Science 359, 97–103.
Iida, N., Dzutsev, A., Stewart, C.A., Smith, L., Bouladoux, N., Weingarten, R.A., Molina, D.A., Salcedo, R., Back, T., Cramer, S., et al. (2013). Commensal bacteria control cancer response to therapy by modulating the tumor microenvironment. Science 342, 967–970.
Laplane, L., Duluc, D., Larmonier, N., Pradeu, T., and Bikfalvi, A. (2018). The Multiple Layers of the Tumor Environment. Trends Cancer 4, 802–809.
Laplane, L., Duluc, D., Bikfalvi, A., Larmonier, N., and Pradeu, T. (2019). Beyond the tumour microenvironment. Int. J. Cancer.
Matson, V., Fessler, J., Bao, R., Chongsuwat, T., Zha, Y., Alegre, M.-L., Luke, J.J., and Gajewski, T.F. (2018). The commensal microbiome is associated with anti–PD-1 efficacy in metastatic melanoma patients. Science 359, 104–108.
Routy, B., Chatelier, E.L., Derosa, L., Duong, C.P.M., Alou, M.T., Daillère, R., Fluckiger, A., Messaoudene, M., Rauber, C., Roberti, M.P., et al. (2018). Gut microbiome influences efficacy of PD-1–based immunotherapy against epithelial tumors. Science 359, 91–97.
Sivan, A., Corrales, L., Hubert, N., Williams, J.B., Aquino-Michaels, K., Earley, Z.M., Benyamin, F.W., Lei, Y.M., Jabri, B., Alegre, M.-L., et al. (2015). Commensal Bifidobacterium promotes antitumor immunity and facilitates anti-PD-L1 efficacy. Science 350, 1084–1089.
Thomas, R.M., and Jobin, C. (2015). The Microbiome and Cancer: Is the ‘Oncobiome’ Mirage Real? Trends Cancer 1, 24–35.
Vétizou, M., Pitt, J.M., Daillère, R., Lepage, P., Waldschmitt, N., Flament, C., Rusakiewicz, S., Routy, B., Roberti, M.P., Duong, C.P.M., et al. (2015). Anticancer immunotherapy by CTLA-4 blockade relies on the gut microbiota. Science 350, 1079–1084.
Viaud, S., Saccheri, F., Mignot, G., Yamazaki, T., Daillère, R., Hannani, D., Enot, D.P., Pfirschke, C., Engblom, C., Pittet, M.J., et al. (2013). The intestinal microbiota modulates the anticancer immune effects of cyclophosphamide. Science 342, 971–976.
Zitvogel, L., Ma, Y., Raoult, D., Kroemer, G., and Gajewski, T.F. (2018). The microbiome in cancer immunotherapy: Diagnostic tools and therapeutic strategies. Science 359, 1366–1370.
 Maya Saleh (ImmunoConcept, UMR5164, CNRS & University of Bordeaux, France & McGill University, Canada)
Maya Saleh (ImmunoConcept, UMR5164, CNRS & University of Bordeaux, France & McGill University, Canada)
Future Directions in Improving Cancer Immunotherapies
Cancer control is intricately linked to the potency and diversity of the immune response. The fundamental understanding of cancer-immune cells crosstalk has resulted in breakthroughs in cancer immunotherapies e.g. by immune checkpoint inhibitors (ICI), among others. However, despite these advances, there is currently an urgent unmet clinical need in oncoimmunology, that is the identification of reproducible predictive biomarkers of response. These include biomarkers that predict 1) efficacy; 2) immune-related adverse events; and 3) hyper-progressive disease, a phenomenon of violent progression of tumor growth in a subset of patients. This is not only crucial for augmenting the proportions of responding patients who achieve durable clinical responses, preventing overtreatment, and tailoring precision medicine with specific ICI (or combinations), but also for the sustainability of emerging and novel cancer immunotherapies. To date, the complexity of the immune response elicited by these immunotherapies in different cancer types and patients has not been well characterized. An integrative analysis that combines in-depth immune phenotyping, functional genomics of the immune and microbial compartments is thus needed towards precision medicine. An improvement in cancer immunotherapies will thus likely require a multi-pronged approach that combines targeting of specific tumor antigens, modulation of the microbiota and epigenetic modifications to elicit a more robust but specific response.
 David Santamaria, (INSERM U1218, IECB, University of Bordeaux, France)
David Santamaria, (INSERM U1218, IECB, University of Bordeaux, France)
Quantitative vs qualitative MAPK signalling in lung adenocarcinoma
The MAPK signalling pathway plays a fundamental role downstream of virtually all known driver oncogenes in lung adenocarcinoma (LUAD). Data from inducible mouse models indicate that its signal intensity determines the nature of the cancer-initiating cell and ultimately the resulting tumour phenotype. In this context, excessive MAPK signalling induces a toxic phenotype causing impaired tumour development that can be rescued by partial MEK inhibition. These evidences suggest that an ideal signalling output might be selected for in advanced tumour stages to maximize disease progression. We have applied a previously described 6-gene transcriptional signature to evaluate the MAPK signalling output in human LUAD. Our observations indicate that the pathway activity is a prognostic factor in KRAS but not in EGFR-mutant patients. Unexpectedly, high MAPK output correlates with extended KRAS-mutant patient survival probably due to excessive signal resulting in replicative stress and slow disease progression. Preliminary evidences indicate that a fraction of these high output patients may display co-existing oncogenic drivers undetected at diagnosis that could contribute to the elevated signalling. We are currently studying this patient cohort as a source to potentially identify novel regulators of the MAPK pathway or potential novel driver events as well as to uncover cellular mechanisms that allow cancer survival in the context of oncogenic toxicity.
 Catherine Sawai (INSERM U1218, Bordeaux, France)
Catherine Sawai (INSERM U1218, Bordeaux, France)
In vivo models of normal and leukemic hematopoiesis
Hematopoietic stem cells (HSCs) differentiate and self-renew, and this activity ensures lifelong hematopoiesis. Nevertheless, several key hematopoietic defects are associated with age, including the increased incidence of myeloid neoplasms. These defects are thought to arise, at least in part, within HSCs, but exactly how this occurs is poorly understood. Experimentally, HSC function is typically measured using transplantation assays, and while this procedure reveals the potential for self-renewal and differentiation, it does not demonstrate HSC function in homeostasis. Unexpectedly, initial reports of HSC function in unperturbed settings suggested that their contribution to steady-state hematopoiesis is minimal. Further studies are therefore required to further resolve the fundamental properties of endogenous HSCs, i.e. in the absence of transplantation, at the steady state. We recently developed transgenic reporters that enable the study of endogenous HSCs. Our system specifically labels the fraction of HSCs with the least differentiated phenotype and the capacity for durable self-renewal. We performed lineage tracing experiments and found that labeled HSCs give rise to other phenotypic HSCs, confirming their position at the top of the hematopoietic hierarchy. In addition to defining the function of HSCs in the steady state, we have begun to investigate the impact of the expression of mutations associated with myeloid malignancy on the endogenous functions of HSCs. Furthermore, building upon our system, we have established a novel model that enables the identification and distinction between normal and leukemic stem cells. This system may be useful in determining the impact of leukemia on normal hematopoiesis as well as studying the functions of leukemic stem cells.
Selected speakers
 Joseph Crompton (Memorial Sloan Kettering Cancer Center, New York, USA)
Joseph Crompton (Memorial Sloan Kettering Cancer Center, New York, USA)
“Cancer” as speech act: the case of papillary thyroid microcarcinoma
It is a commonly held view in medicine that diagnostic terms faithfully describe the reality of the biological phenomena to which they refer. Here I raise the possibility that the diagnostic term “cancer” does not always denote a bona-fide malignancy, but can function as speech act, transforming the medical reality of the biological phenomena it is meant to identify and describe. As a speech act, “cancer” performs an action, prompting cancer doctors and patients to undergo the same diagnostic and therapeutic approaches that are used for the diagnosis and treatment of true malignancies.
Locutionary
As a speech act with performativity, “cancer” has locutionary, illocutionary, and perlocutionary functions. For both the public/patients, and medical professionals, the locutionary term “cancer” cannot be perceived separate and apart from its innumerable “piggy-backing” meanings and associations. Formal definitions and diagnostic criteria can be vague and ambiguous, setting up the risk that the term “cancer” can denote lesions with malignant features, but are not true malignancies. The problem is most clearly seen with lesions that grossly or histologically resemble true malignancies or demonstrate hallmark malignant behavior such as lymph node metastases, but for which the natural history is incompletely understood. Three examples of common lesions that are designated as “cancers”, but have an uncertain malignant potential, are (i) papillary thyroid microcarcinoma, (ii) low-grade ductal carcinoma in situ of the breast, and (iii) low-grade prostate cancer.
Papillary thyroid microcarcinoma is an entity that was described as early as 19271, but whose incidence has skyrocketed in the last thirty years. The initial identification of papillary thyroid microcarcinoma came from thyroid specimens, which were not resected for true malignancy, but for other indications, such as benign multinodular goiter. Further characterization of these lesions with autopsy studies showed that less than 10% of people who died from non-cancerous etiologies harbored papillary thyroid microcarcinoma, suggesting it may be a tumor that does not cause symptoms or death the way a true malignancy would if left untreated.
Recognizing the benignancy of these lesions, pathologists initially cautioned against the use of the word “carcinoma” in describing them. In a report in 1949, Hazard et al2 note “It is important that the innocence of the nonencapsulated sclerosing tumor of the thyroid [former name for papillary thyroid microcarcinoma] be recognized and appreciated…the surgeon may become unduly alarmed when the pathologist reports the presence of carcinoma. This may lead to reoperation, radical dissection of the neck or extensive irradiation, all of which are unnecessary and undesirable. If the true nature of the tumor is recognized its presence can be disregarded.” The authors had an intuitive sense that the word “carcinoma” comes with significant consequences for patients. It is interesting to note this preceded the publication of How to Do Things with Words, J.L. Austin’s most influential work on speech acts. (Incidentally, Austin died of lung cancer at age 48).
Several years later in 1955, Klinck et al3 published a study arguing that papillary thyroid microcarcinoma should be considered a true malignancy because, in some cases, it can metastasize to lymph nodes. Although they did not show these lesions cause significant symptoms or death, they concluded in their paper: “the term nonencapsulated sclerosing tumor is misleading and should not be used. To replace it, we suggest the term occult sclerosing thyroid carcinoma.” The use of the word “carcinoma” by these authors galvanized its identity as a true malignancy and it has, thereafter, been treated as such. The worry here is that, in calling these lesions “carcinoma”, the authors uttered a term with locutionary performativity that has profoundly altered the way these lesions treated, even though our understanding of them remains limited.
Illocutionary
The illocutionary act of “cancer” is the active result of the meaning implied by the locutionary sense; in naming these lesions “cancer”, the term effectively renders them malignant in a medical sense (but not necessarily in a biologic sense). In the case of papillary thyroid microcarcioma, the speech act of “cancer” results in the transformation of a person with a potentially benign lesion into a cancer patient.
An additional problem for the medical field is that, once a lesion is designated as a “cancer”, a better understanding of its natural history is difficult to elucidate because we intervene with medical treatments, such as surgery. The possibility that papillary thyroid microcarcinoma can grow, spread, cause symptoms and death is confounded by removing the thyroid—it may be that it has true malignant potential, but it is also possible that the lesion never comes to clinical attention.
In the case of papillary thyroid microcarcinoma, because it is almost always asymptomatic and cannot be palpated on physical exam, the incidence has historically been low. In the 1990s, however, increased screening with ultrasound and fine-needle aspiration biopsies spurred a pandemic of papillary thyroid microcarcinoma; treatment with partial or complete thyroidectomy became the standard of care4. In South Korea, for example, the incidence increased more than sevenfold between1999 to 2009—from 6.3 per 100,000 to 47.4 per 100,000 people. Curiously, mortality from thyroid cancer has remained unchanged, suggesting either: (i) treatment is nearly always effective (rare for cancer treatments), or (ii) papillary thyroid microcarcinoma has an indolent natural history that may not require intervention.
Even while partial or complete thyroidectomy became the standard of care for papillary thyroid microcarcinoma across the globe, a physician in Japan, Akira Miyauchi MD, PhD, opted to, instead, closely surveil his patients with papillary thyroid microcarcinoma. Rather than use a thyroid ultrasound to diagnose and surgically remove tumors, he used it to closely observe the lesions and better understand their natural history. If any patient demonstrated growth of the tumor or spread to lymph nodes, he would remove them surgically. Remarkably, with over 20 years follow-up, very few patients have required operative intervention.
Perlocutionary
The main consequence of a “cancer” speech act is that the locutionary and illocutionary forces result in the perlocutionary effect of cancer treatment: surgery, radiation, and chemotherapy are treatments with serious adverse effects and unclear therapeutic benefit for potentially benign lesions. In the case of papillary thyroid microcarcinoma, thyroidectomy comes with significant short-term risks of bleeding and infection. The long-term risks of surgery for some patients include damage to the recurrent laryngeal nerve, causing changes in the voice and even difficulty breathing. All patients with total thyroidectomy will require life-long thyroid hormone replacement. This is to say, speech acts potentially come with great cost to patients for unclear benefit. The larger concern here is the possibility that the current therapeutic approach to a handful of tumors is a colossal speech act, collectively articulated by the medical field, and executed by individual practitioners; that is, the locutionary act of naming a potential benignancy “cancer” has the illocutionary effect of it becoming cancer and the perlocutionary consequence of it being treated like cancer.
References
Furuya-Kanamori et al. Prevalence of Differentiated Thyroid Cancer in Autopsy Studies over Six Decades: A meta-analysis. Journal of Clinical Oncology. 2016.
Hazard et al. Nonencapsulated Sclerosing Tumors of the Thyroid. J Clin Endocrinol Metab. 1949.
Klinck et al. Occult Sclerosing Carcinoma of the Thyroid. Cancer. 1955.
Ahn et al. Korea’s Thyroid-Cancer “Epidemic”—screening and overdiagnosis. New England Journal of Medicine. 2014.
 Gregor Greslehner (ERC IDEM, CNRS, Bordeaux, France)
Gregor Greslehner (ERC IDEM, CNRS, Bordeaux, France)
Explaining the common biology of aging and cancer development
The biological phenomena of aging and cancer development are surrounded by several conceptual questions. Since it affects all individuals of multicellular species, it can be asked whether aging should be viewed as a disease. Indeed, debating this very question has recently become a hot topic in biology and philosophy of medicine. In any event, there are a number of age-associated diseases. Age is one of the main risk factors for several diseases, including most prominently cancer. Since the population continues to grow older, this has huge implications for society. If aging were a disease, can it be cured? If it cannot be cured, can the aging process at least be slowed down? Evidence suggests that there is a maximum lifespan for human being around 120 years. Cancer and aging can be seen at both ends of a spectrum which keeps dividing cells and bay and which does not eliminate cells too easily that are needed to keep the tissue intact. There are a number of hallmarks of cancer (Hanahan and Weinberg, 2011) and aging (López-Otín et al., 2013)—some of which both have in common (Aunan et al., 2017; Finkel et al., 2007).
Cancer and aging can be arguably considered to constitute two sides of the same coin: on the one hand, dividing cells need to be kept under control and arrested in a state of senescence once they have acquired certain defects, on the other hand, senescent and non-dividing cells lead to decreased cell numbers in tissues, resulting in the age-associated loss of function in these tissues, be it in the skin, hair, bones, brain, etc. The mechanism responsible for this is cellular senescence, a key player in both aging and cancer. Failure in finding the right balance can lead to either cancer or aging phenotypes.
No individual hallmark is sufficient to explain why multicellular organisms age. Several molecular and cellular factors have been identified to conjunctively being responsible for the phenomenon of aging in multicellular organisms. On the other hand, satisfactory explanations of aging need to go further than the underlying physiological mechanisms. When it comes to questions of the evolutionary dynamics of aging and differences in lifespan, one needs to look beyond the individual and address questions concerning natural selection and its role in bringing about organisms having different lifespans. There exists a multitude of both physiological and evolutionary explanations about the phenomenon of aging. Z. A. Medvedev systematized more than 300 theories of aging in “an attempt at a rational classification of theories of aging” (Medvedev, 1990). That is an impressive multitude of explanatory narratives for a biological phenomenon. Perhaps the most striking difference in explaining aging is whether one addresses either the ontogenetic or phylogenetic aspects of aging, i.e., (i) the particular molecular or physiological mechanisms, or (ii) the evolutionary aspects of longevity, like for example the optimal rate of total lifespan and reproductive period.
Taken together, aging and cancer research show a general lesson for current biological practice: the need to combine different explanatory accounts for dealing with complex biological phenomena, which are being addressed by a multitude of different scientific disciplines. Viewed this way, one can conjecture that molecular mechanistic explanations and systems biology explanations are not alternatives, but rather complementary at dealing with different aspects of biological phenomena at different levels of abstraction. Given the interdisciplinary tradition within the life sciences, however, this might be a debate about labels rather than about actual research practice.
While it can be debated whether or not aging should be considered a disease, the fact that cancer is one usually goes without question. However, the next step is to further classify different forms of cancer. Cancer is not a single disease. Therefore, classification is an essential starting point for cancer diagnosis and treatment. Various classification systems exist in medical and biological practice. The dominant approach is to classify cancer based on the cellular type of its origin (Hoadley et al., 2018). Recently, however, other molecular classification systems have been developed which allow for more fine-grained classification—and more targeted therapies. While the tissue from which cancer originates is an important factor, key molecules are shared in cancer types from different tissues and different types of cancer can arise within the same tissue.
Each type of cancer begins with a single cell hidden in its adjacent tissue for a long time. Once cancer becomes detectable clinically, there usually need to be more than a billion cancer cells. Being a slow process, cancer can remain below the threshold of detectability for the major part of its genesis. For early detection, certain biomarkers like DNA mutations and proteins present in the blood are an important diagnostic tool. The importance of other cellular and physiological properties for classifying and treating cancer is being stressed, based on recent scientific and medical practice. This way, a much more fine-grained molecular characterization and classification of cancer becomes possible. If certain classes of target protein domains can be identified, this also suggests ways to arm the immune system against these protein domains already at early stages of cancer.
A better molecular understanding of the structural and functional properties of affected protein in cancer cells is the first step to understanding the processes on higher levels of organization. Questions that need to be addressed include: Which hallmark protein domains are affected in which types of cancer? Which structural and functional properties of such domains cause cells to become malignant? Does protein-based classification deviate from cell-of-origin classification? The key proteins appear to be involved in regulating the balance between keeping neoplastic cells at bay and not overcautiously stopping cells from proliferating without need. Apoptosis and cellular senescence, i.e. cells killing themselves or arresting themselves in a state of non-division, are triggered by certain core proteins (tumor suppressors). Frequently, it is exactly such proteins which are affected in different types of cancer. Adequate accounts for classifying and explaining cancer need to address these questions.
Like for aging, several explanatory accounts have been proposed that need to be combined in order to explain the phenomena and processes of cancer, including mechanistic and evolutionary accounts (Lean and Plutynski, 2016).
References
Aunan, J. R., et al. (2017). The biology of aging and cancer: A brief overview of shared and divergent molecular hallmarks. Aging and Disease, 8(5):628-642. doi:10.14336/AD.2017.0103.
Finkel, T., et al. (2007). The common biology of cancer and ageing. Nature, 448:767-774. doi:10.1038/nature05985.
Hanahan, D. and Weinberg, R. A. (2011). Hallmarks of cancer: The next generation. Cell, 144(5):646-674. doi:10.1016/j.cell.2011.02.013.
Hoadley, K. A., et al. (2018). Cell-of-origin patterns dominate the molecular classification of 10,000 tumors from 33 types of cancer. Cell, 173(5):291-304. doi:10.1016/j.cell.2018.03.022.
Lean, C. and Plutynski, A. (2016). The evolution of failure: explaining cancer as an evolutionary process. Biology & Philosophy, 31:39-57. doi:10.1007/s10539-015-9511-1.
López-OtÍn, et al. (2013). The hallmarks of aging. Cell, 153:1194-1217. doi:10.1016/j.cell.2013.05.039.
Medvedev, Z. A. (1990). An attempt to a rational classification of theories of aging. Biological Reviews of the Cambridge Philosophical Society, 65(3)
 Laurence Huc (INRA, Toulouse, France)
Laurence Huc (INRA, Toulouse, France)
What is a carcinogen? bridging the gap between basic biology and regulatory toxicology
The origins of cancer is a central question for biologists. Even if Rachel Carson established the link between chemical contaminated-environment and cancer in the 1960’s, the hegemonic area of genetics gave the direction of research in oncology and carcinogenesis. From “back luck” origins to the genetic causes of cancer, the part of environment in the occurrence of this disease is a central point of debate regarding public health. What does it make controversy? Why is it so hard to define a carcinogen?
Through current events of controversies, with the example of persistent pollutants like dioxin and hydrocarbons and with the example of pesticides such as glyphosate, I will illustrate the definitions of a carcinogen, according to the point of view of a biologist, a toxicologist in basic science. I will also focus on the definition supported by the non-profit organization IARC (International Agency for Research on Cancer) and the marked discrepancy with the regulatory agencies like EFSA (European Food Safety Authority) and ECHA (European Chemicals agency). Then, I will examine the proofs of carcinogenicity according to laws and citizens. Finally, I will analyse the influences of the private interests and think tanks on the scientific definition of a carcinogen and how science is deprived of its own produced knowledge.
 Andrea Pensotti (University of Rome, University Campus Bio-Medico of Rome, Italy), Marriano Bizzarri (Sapienza University, Italy),
Andrea Pensotti (University of Rome, University Campus Bio-Medico of Rome, Italy), Marriano Bizzarri (Sapienza University, Italy),
and Marta Bertolaso (University of Rome, University Campus Bio-Medico of Rome, Italy)
Epistemic premises pave the way of scientific methodology in cancer studies: tumor reversion as case study
“How do we know that we know?” Scientific knowledge cannot be viewed as an absolute, but is actually constrained by the theoretical premises on which experiments are planned and the subsequent acquired raw data are further interpreted. Ultimately, there is currently a wide agreement on the fact that Science is a way of knowing that requires a strong philosophical underpinning (whether consciously sought of unconsciously learned), which could provide an operational and reliable definition of some key conceptual tools such as causality or system, on which scientific practice is forced to deal with.
Every theory has thus a relative character and should be viewed as an asymptotic attempt to grasp the real word (a model is still a fictional representation as epitomized by Magritte’s picture). Yet, we suggest that the reliability of an epistemological framework lies on the number of phenomena that this theory can actually explain and on how many previous paradoxes can be solved with the help of this new approach.
A clear-cut example is provided by studies highlighting the phenotypic reversion in a number of cancer types. Recent experiments performed in our lab, clearly show that both natural compounds (like inositol) or microRNA extracted from embryonic micro-environment, can effectively induce tumor reversion through the involvement of a few critical pathways, including: activation of p53, redistribution of E-cadherin/β-catenin, cytoskeleton remodeling, down regulation of TCTP expression, just to mention a few. Overall, those data suggest that the tumor reversion involves a specific “molecular programme”, which can be triggered by soluble molecular factors as well as by biophysical cues.
Tumor reversion processes were observed since the first decade of the XX century, but the reductionist approach – on which biology mostly relies – was unable to accommodate such findings within the current accepted frameworks. Therefore for almost a century, scientists developed technology and experiments almost only focused on the search of “key targets”, able in identifying cancer cells and hence kill them. The emergence of the theory of the complexity – which undermines the scale-free, linear assumptions underpinning reductionism – and the accumulation of an increasing burden of phenomena that cannot be explained by the current theoretical paradigm, prompted a number of scholars in reconsidering the philosophical basis, i.e. the epistemological premises on which cancer research is grounded.
A more comprehensive theory of carcinogenesis should in principle be able to deal with complexities as well as with the overwhelming body of paradoxes gathered until now (tumor reversion induced by morphogenetic factors, questionable role of mutations, non-genotoxic carcinogenesis, just to mention a few), with which the current paradigm can hardly accommodate. Taming this complexity does not means we should manage more data (as we are already submerged by an intractable huge body of information, often irrelevant, produced by the use, or abuse of technologies). Instead, we should be able in extracting a few set of data and of relevant relationships that could help in grasping the dynamics of the process under scrutiny. Namely, Systems Biology allows to think biological systems not as – complex as they can be – organized molecular systems, but mostly as organizers of living organisms. This would not only answer old questions (and inexplicable paradoxes), but will drive questions over the dynamic transitions occurring in living systems towards health or disease.
Choosing between competing premises and testing alternative theoretical hypothesis have been the core component of the experimental science since the Renaissance. However, a widely accepted paradigm will hardly be dropped before a considerable amount of paradoxes and contradictions has been resolved. Such moment seems to have come. For a while, cancer has been considered like a Spinoza’s monad, that is, as a microcosm emerging from a deregulated genome occurring in a single cell. Yet, that paradigm has encompassed several shortfalls and it is now increasingly recognized that the overall framework is very much complex that previously thought. We have to regard carcinogenesis and carcinomas as errors in development occurring in tissues, not in individual cancer cells, as the experimental setup we focus on for tumor reversion shows. As such, those “phenotypes gone awry” should not be viewed as irreversibly committed: instead, even wrong phenotypes can be “reprogrammed” and committed toward a normal differentiating pathway. Both soluble factors (mRNAs, inositol, etc.) and biophysical cues have showed to promote cancer reversion, by displaying a number of pleiotropic effects through the involvement of several targets. This approach can open new affordable avenues for therapeutic solutions. The successful results obtained in treating even few selected cancers through microenvironmentdrug modifiers, have firmly established the paradigm of “differentiation therapy” as a valid approach for the reversion of malignancy in the treatment of neoplasia in human patients. Further studies should be pursued by combining the use of well-designed pharmacological substances, as well as nutrients and other natural compounds, able to hit microenvironment-based targets. Finally this approach introduce not only a new framework for experiments design, but also ask for a new regulatory approach in order to include this new “systemic treatments” which are neither comparable to a pharmacological model, nor to a food supplement model. Therefore, reshaping the theoretical approach in biology of cancer means also to rethink the way clinicians and institutions responsible for public health are addressing cancer management and study. Along this perspective, it is also likely that new opportunities for pharma industry would be disclosed, given that companies have the opportunity to translate this theoretical approach in new business models, “epistemically driven”.
References:
Bissell MJ, Radisky DC, Rizki A, Weaver VM, Petersen OW (2002) The organizing principle microenvironmental influences in the normal and malignant breast. Differentiation 70:537–546
Hendrix MJC, Seftor EA, Seftor REB, Kasemeier-Kulesa J, Kulesa PM, Postovit L-M (2007) Reprogramming metastatic tumour cells with embryonic microenvironments. Nat Rev Cancer 7:246–255
Wang F, Hansen RK, Radisky D, Yoend T, Barcellos-Hoff MH, Petersen OW, Turley EA, Bissell MJ (2002) Phenotypic reversion or death of cancer cells by altering signalling pathways in three- dimensional contexts. J Natl Cancer Inst 94:1494–1503
White FM, Gatenby RA, Fischbach C. (2019) The Physics of Cancer, Cancer Res. 2019 May 1;79(9):2107-2110
Amson R1, Karp JE, Telerman A., Lessons from tumor reversion for cancer treatment. Curr Opin Oncol. 2013 Jan;25(1):59-65
Amson R1, Pece S, Marine JC, Di Fiore PP, Telerman A. TPT1/ TCTP-regulated pathways in phenotypic reprogramming. Trends Cell Biol. 2013 Jan;23(1):37-46
Huang, S., Ingber, D.E., 2007. A non-genetic basis for cancer progression and metastasis: self-organizing attractors in cell regulatory networks. Breast Dis. 26, 27e54.
 Elena Rondeau (University of Bordeaux, France)
Elena Rondeau (University of Bordeaux, France)
Understanding cancer progression and its control: An analysis of immune contribution to metastasis
Ongoing progress in cancer research is improving our comprehension of the spatial and temporal factors influencing tumour progression, indeed characterised by the pleiotropic involvement of multiple cellular and molecular elements present in the tissue environment at different stages of the disease (Plutynski 2018). While supplementing our thorough appreciation of the cell-intrinsic properties of transformation, these advances also illustrate a noticeable evolution in our investigation of cancer causality, i.e. enriching cell –centered research programs with the acknowledgement of the crucial contribution of environmental factors at various levels, from the tumour milieu (or “micro-environment”) to the host “macro-environment” (Al-Zoughbi 2019). Despite the significance of this conceptual shift, scientists and clinicians alike still face serious difficulties, in particular those linked to tumour recurrence and distant spread.
In this talk, I explore some conceptual issues linked to our current understanding of cancer progression at the tissue- and organism- levels, with a particular focus on the systemic dimension of metastasis and the involvement of host immunity.
Tackling the complexity of organ-specific dissemination:
Metastasis is currently the leading cause of cancer mortality, correlating with disease severity and resistance to conventional therapy. The accepted definition of this multistep process describes it as a succession of events through which tumour cells exit the primary mass and invade surrounding tissues, then enter the circulation and migrate to distant implantation sites, where they develop as secondary growths.
However, such characteristics of linearity and sequentiality are now being questioned by the existence of feedback loops and retrograde interactions between the different actors and steps of metastasis. This sets a context of causal complexity and intricacy between cancer cell –intrinsic and –extrinsic factors, where several key observations remain partly unexplained and conceptually challenging. One interesting example is the report of organ specificity for secondary growth, which accounts for the apparent tropism of migrating cells toward certain metastatic tissues. Since the beginning of cancer research and medicine, oncologists have indeed been struck by the statistical correspondence between the nature of the primary tumour and the location of secondary growths (for instance, breast tumours usually spread to the bone or lung, whereas colorectal cancer often leads to liver metastasis).
A historical perspective on metastasis research:
The “seed and soil” theory of metastasis, initially coined by British surgeon Stephen Paget (1889), stipulates that such patterns can be explained by favourable interactions between circulating tumour cells (the “seed”) and specific microenvironments encountered during their migration (the “soil”). Initially based on autopsy studies of the affected organs in cancerous patients, this hypothesis was soon replaced with a mechanical explanation for tumour cell arrest (Ewing 1929), supposedly determined by the anatomy of vascular connections between the primary site and further organs accessible through blood circulation. Metastasis research in the early twentieth century adopted the latter proposal in its experimental designs, before the discovery of cancer cell heterogeneity and the refutation of mechanical exclusiveness in the causality of dissemination. Paget’s forgotten analogy then resurfaced as an appropriate and testable explanation for organ selectivity in secondary tumour growth (Fidler 2003).
It therefore seems interesting to examine the historical dynamics of this debate and its conceptual implications, which may directly participate to shaping current research hypotheses concerning the yet unanswered question of exactly “what decides which organs suffer in disseminated cancer?” (Paget 1889). With this challenge of better understanding the connection between primary and metastatic tumours, comes an incentive to re-evaluate the relative efficiency and spatio-temporal compatibility between the phenomena of circulation, colonisation and secondary organ attractiveness (as currently discussed in the field, such as Lu 2019).
Aiming to analyse and revisit the “seed and soil” analogy:
Starting with an effort to contextualise Paget’s claims on metastasis causality, the objective here is to better characterise their epistemological status and heritage, as the “seed and soil” analogy has been, and presently is, extensively cited and re-interpreted in the scientific literature. Given this perspective, the premises and consequences of the theory could, beyond maintaining their illustrative role, benefit from a revisited analysis in the light of recent clinical and experimental observations. In particular, the evidence of distant “pre-metastatic niche” preparation reveals anticipatory signalling in the process of cancer dissemination (Chin 2016). It corresponds to the establishment of a tumour-promoting milieu in the secondary organ, before the arrival of cancer cells. This phenomenon involves both tumour-derived factors and pre-existing host components hijacked by the malignant context, thus refining our consideration of secondary organ specificity.
Crucial in this reappraisal is the aim to decipher the implication of the immune system in cancer progression, given the delicate balance between its protective and pro-tumoral activities. Indeed pre-metastatic processes are frequently associated with immune cell recruitment, inflammatory signalling and structural remodelling, with a direct contribution of these features to tumour development and further seeding. As a matter of fact, some populations of suppressive and pro-invasive immune cells may actively foster “seed” persistence and enable distant metastasis by both establishing a hospitable or attractive environment in future “soils” and clustering with circulating cancer cells.
Characterisation of immune involvement in organ tropism:
Hence the pressing question of determining the exact role of host immunity in metastasis causality: to what extent, and how, do the immune actors of cancer progression contribute to the preparation and specificity of secondary sites? In what ways do they differentially behave and interact according to the spatio-temporal context? Underlying the conceptual framework of the “seed and soil” analogy, would this reflection help better understand the relative contributions of pro-metastatic factors, and how may it affect practical and therapeutic strategies?
These interrogations also uncover an important consideration on our vision of cancer progression, namely the impact of when and where to focus when studying the disease. It may indeed prove useful to direct investigations on the pre-existing and early-stage properties of host components that are later involved in metastasis (especially when considering the immune landscape), as well as on the characteristics of unaffected tissues, in order to clarify the arguments of secondary organ predisposition and/or diminished resistance to tumour implantation.
In my research, I propose to address this issue from both a conceptual and an experimental point of view, through a case study of metastatic tropism in a mouse model of triple negative breast cancer, known to specifically disseminate to the lungs: could we define a phenomenon of immune “fertilisation” here?
Rationale of the MDSC case study:
In our model of induced tumorigenesis (4T1 cells injected directly into the mammary gland), cancer progression correlates with an early and gradual accumulation of a subset of immature immune cells (named “myeloid-derived suppressor cells”, MDSCs) specifically in the spleen and in the lung (pre)-metastatic niche. These MDSCs can be categorised into two subpopulations (granulocytic and monocytic), which both participate to cancer-promoting inflammation and metastasis, through their immunosuppressive, pro-angiogenic and pro-migratory activities (Wang 2019). In an attempt to better understand their disease-related skewing and their role in lung tropism for secondary tumour growth, the characteristics of these myeloid cells are monitored through the course of tumour evolution and in differential environments, including primary tumour, pulmonary tissue, non-metastatic liver, lymphoid organs and blood.
Using an appropriate combination of surface markers, we have observed a novel and dynamic profile of MDSC presence in the lungs, as the evidence for three subpopulations may complement the aforementioned dual view. These subsets each see their percentages vary through the course of cancer progression, with an interesting shift between the relative importance of the two granulocytic groups, which may reveal differential properties according to stage and location. Current works aim to refine this MDSC phenotyping while tracking the tumour cells as they spread to the lungs (in vivo imaging with marked 4T1 cells), as well as exploring the functional characteristics of these myeloid populations and their plasticity.
As a whole, this interdisciplinary project aims to address the current challenge of better understanding organ-specific metastasis, through a dialogue between conceptual analysis and experimental illustration. Studying the heritage and update of the “seed and soil” analogy uncovers relevant interrogations about the spatio-temporal dynamics of cancer dissemination and the contribution of host factors in early determination of secondary tumour seeding.
References:
Al-Zoughbi, W., & Hoefler, G. (2019). Tumor Macroenvironment: An Update. Pathobiology, 1-3.
Chin, A.R., and Wang, S.E. (2016). Cancer Tills the Premetastatic Field: Mechanistic Basis and Clinical Implications. Clin Cancer Res 22, 3725–3733.
Ewing, J. Neoplastic Diseases edn 6 (W. B. Saunders, Philadelphia, 1928).
Fidler IJ. The pathogenesis of cancer metastasis: the `seed and soil’ hypothesis revisited. Nat Rev Cancer. 2003; 3:453–8.
Lu, Y., Lian, S., Cheng, Y., Ye, Y., Xie, X., Fu, C., … & Jia, L. (2019). Circulation patterns and seed-soil compatibility factors cooperate to cause cancer organ-specific metastasis. Experimental cell research, 375(1), 62-72.
Paget, S. The distribution of secondary growths in cancer of the breast. Lancet 1, 571–573 (1889).
Plutynski, A. (2018). Explaining cancer: Finding order in disorder. Oxford University Press.
Wang, Y., Ding, Y., Guo, N., & Wang, S. (2019). MDSCs: Key Criminals of Tumor Pre-metastatic Niche Formation. Frontiers in Immunology, 10.
 Lucas Serra (University at Buffalo, USA)
Lucas Serra (University at Buffalo, USA)
Ontological Considerations in the Representation of Cancer
The National Cancer Institute spends nearly 5 billion dollars annually on cancer research a year. This work, alongside privately funded ventures, produces incomprehensible mountains of data each year regarding cancer biology, pathology, staging, treatments, and outcomes. In order to manage these mountains, a plethora of databases, classification systems, and ontologies have been developed. This paper addresses numerous ontological considerations such as: What is the essence of cancer? How do we represent cancer and its associated entities in a logically coherent manner in an ontology? What prior work has been done within this field? Lastly, we examine our own work in representing hematologic malignancies within an ontology and the challenges faced in its implementation.
The field of ontology lies at a crossroads between philosophy and computer science. In philosophy, ontology is the study of being and is a branch of metaphysics that has existed since the time of early Greek philosophers like Parmenides and Aristotle. (1) A more recent notion of ontology, derived from the field of computer science, regards ontologies as reusable libraries of knowledge that are created under specific design principles and methodologies. (2) For the purposes of this paper we will use the following definition of an ontology: “A representational artifact, comprising a taxonomy as proper part, whose representations are intended to designate some combination of universals, defined classes, and certain relations between them.” (3) Essentially, ontologies are taxonomies that seek to faithfully represent entities within our world. Most of the commonly used ontologies of today are created from an ontological realist perspective. (4)
The primary purpose of an ontology is to enhance the interoperability of data by standardizing terms and providing a semantic framework. To this end, ontologies include human and machine-readable definitions for entities within its hierarchy. Human-readable definitions are comprised of text, ideally, in the form of an “Aristotelean definition”, which includes a reference to the term’s parent class along with a statement about the differentiae. The differentiae is a property that demarcates the term from other child terms assigned to the parent class. (5) Machine-readable definitions take the form of formal, logical axioms and help to unambiguously describe an entity.
With these ontological ideas in mind, we can now begin to ask: What is the essence of cancer? How should we go about representing it within an ontology? There are multiple levels to dissect when approaching such a broad and loaded topic such as cancer. One of the most important steps is to distinguish cancer (the disease) against cancer (as it exists as tumors) and cancer (the disease course). Some thought has already been given to this issue in a paper that gave rise to the Ontology for General Medical Science (OGMS). (6) Here, diseases are “dispositions” or attributes of an organism that will initiate specific processes when specific conditions are satisfied. The disease disposition to undergo pathological processes exists due to one or more disorders in an organism. Disorders are “a causally relatively isolated combination of physical components that is (a) clinically abnormal and (b) maximal, in the sense that it is not a part of some larger such combination” and root the disease disposition in a physical basis. The disease course on the other hand refers to the totality of the aforementioned pathological process wherein the disease is realized. Under this view, the authors posit that although individual cells can be disordered, they regard the tumor as the disorder as it the maximal collection of these disordered cells. This view is limited and does not align with recent work regarding the importance of intra-tumoral heterogeneity. (7) An argument could be made that sub-populations of cells within a tumor are “casually isolated”, being derived from the same sub-clone, and share more defining traits, like treatment resistance, than neighboring sub-populations. In essence, the OGMS definition of disorder ignores important functional considerations of these physical components by being overly concerned with the structural unity of these entities.
Later ontological efforts used the framework built by OGMS. The Disease Ontology (DO) represents over 8000 inherited, developmental, and acquired human diseases and seeks to semantically integrate a wide variety of medical terminologies and ontologies through cross mapping. (8) Although the DO adheres to the OGMS ideas of disease, dispositions, and disorders, it occasionally conflates terms. For instance, the DO incorrectly classifies a neoplasm (the physical entity) as a cancer (the disease disposition) in its “ovarian cancer” branch. (9) The the Cell Ontology (CL), which attempts to catalog all human cell types, does not cover entities like diseases but proposes the following definition of a neoplastic cell: “an abnormal cell exhibiting dysregulation of cell proliferation or programmed cell death and capable of forming a neoplasm”. Malignant cells are subsequently defined as neoplastic cells that can invade surrounding tissue. (10) This definition is much closer to capturing the necessary traits that define cancerous cells. Ideally, such a definition would include widely agreed upon features like the so-called “hallmarks of cancer”, which include inducing angiogenesis, replicative immortality, and newly elucidated interactions between tumor cells and the microenvironment. (11)
In our own work, we have created an ontology, the Cancer Cell Ontology, to represent hematologic malignancies, namely acute lymphoblastic leukemia (ALL) and acute myeloblastic leukemia (AML). (12) Our ontology contains 300 terms with human and machine-readable definitions representing cancer cells found in blood cancers and were created following conventions begun by the Cell Ontology. Due to the types of data available to us, we defined cancer cell types solely by their immunophenotypes or surface markers. This highlights an issue in the ontological representation of cancer: completeness. Cancer is an immensely diverse topic with numerous avenues of diagnosis and the usual work-up of a patient could include complete blood counts, bone marrow biopsies, flow cytometry, cytogenetics, biomarkers, and imaging, to name just a few. To capture all of these facets would be extremely difficult within one ontology. There is a careful balance between achieving ontological purity by creating faithful, highly detailed representations of reality and the usability of these constructs. A second ontological issue with the work concerns what we accept as ground truth. One lab may claim that one set of markers identifies AML while another lab may suggest a slightly different antibody panel. This is the case in the United States where antibody panels for blood cancers are not entirely standardized. Who to believe? One strategy is to look for commonalities between identifiers and accept those as truth. Fortunately, in Europe, a group of labs belonging to the Euroflow consortium have reached consensus upon standardized panels, and serve as the basis for the classifications within Cancer Cell Ontology. A related issue affects what entities constitute biomarkers. A recent paper rejects the notion that an entity becomes a biomarker “after and because” it has been measured and instead posits that an entity is a biomarker due to intrinsic characteristics like observability and measurability. (13) This is a sound approach that focuses on the essence of these entities rather than their identity after their use.
In summation, numerous past efforts have examined the ontological basis of disease and cancer. Our representations and ideas of cancer need to evolve and include the latest research. The titanic scope of the cancer research field along with occasionally conflicting perspectives on the nature of diagnostic entities can stymy our efforts in the ontological representation of malignancies.
References
1. Cohen SM. Aristotle’s Metaphysics. https://plato.stanford.edu/archives/win2016/entries/aristotle-metaphysics/: Metaphysics Research Lab, Stanford University; 2016.
2. Gruber TR. Toward principles for the design of ontologies used for knowledge sharing? International Journal of Human-Computer Studies. 1995;43(5):907-28.
3. Arp R, Smith B, Spear AD. Building Ontologies with Basic Formal Ontology: The MIT Press; 2015. 248 p.
4. Smith B, Ceusters W. Ontological realism: A methodology for coordinated evolution of scientific ontologies. Appl Ontol. 2010;5(3-4):139-88.
5. Rosse C, Mejino JLV. A reference ontology for biomedical informatics: the Foundational Model of Anatomy. Journal of Biomedical Informatics. 2003;36(6):478-500.
6. Scheuermann RH, Ceusters W, Smith B. Toward an ontological treatment of disease and diagnosis. Summit Transl Bioinform. 2009;2009:116-20.
7. Liu J, Dang H, Wang XW. The significance of intertumor and intratumor heterogeneity in liver cancer. Experimental & Molecular Medicine. 2018;50(1):e416-e.
8. Schriml LM, Arze C, Nadendla S, Chang Y-WW, Mazaitis M, Felix V, et al. Disease Ontology: a backbone for disease semantic integration. Nucleic Acids Res. 2012;40(Database issue):D940-D6.
9. Duncan WD, Gaudioso C, Diehl AD. Towards an Ontology for Representing Malignant Neoplasms. ICBO Conference Proceedings. 2017.
10. Meehan TF, Masci AM, Abdulla A, Cowell LG, Blake JA, Mungall CJ, et al. Logical Development of the Cell Ontology. BMC Bioinformatics. 2011;12(1):6.
11. Hanahan D, Weinberg Robert A. Hallmarks of Cancer: The Next Generation. Cell. 2011;144(5):646-74.
12. [Deleted for review]
13. Ceusters W, Smith B. Biomarkers in the ontology for general medical science. Studies in health technology and informatics. 2015;210:155-9.
 Michael J. Young (Harvard Medical School, USA)
Michael J. Young (Harvard Medical School, USA)
Epistemic Upshots of the Genomic Turn in Cancer Care: Lessons from Neuro-Oncology
The meteoric rise of genomic assays to stratify, diagnose and understand tumor typology and development has dramatically impacted how cancer is researched, diagnosed, prognosticated and treated. In this presentation, after explaining and contextualizing these recent advances, I evaluate the epistemic and ontologic upshots of this genomic turn in cancer research and clinical care, particularly through the lense of the recently evolving landscape of neuro-oncology. I argue that despite the prima facie promise of mitigating vagueness across boundaries of diagnostic categories, the introduction of genomic properties into nosologic schemata introduces an array of underexplored epistemic challenges in how to ascertain the bounderies of oncologic disease and in how and when to communicate diagnoses to patients.
The talk begins with an overview of recent transformation of the neuro-oncology landscape from a classical histological diagnostic approach to the newly revised genomic approach as encapsulated in the 2016 World Health Organization classification system. In 2016, the World Health Organization (WHO) undertook a dramatic restructuring of its classification of central nervous system (CNS) tumors (i.e., tumors of the brain, spinal cord and nerves), incorporating genetic criteria for the characterization of many tumor types. For example, the revised schema requires characterization of isocitrate dehydrogenase (IDH) mutation status as part of the histologic classification of gliomas, and requires 1p19q codeletion and IDH mutation for the classification of oligodendrogliomas. The integration of histologic and molecular parameters into the revised nosology for central nervous system tumors reflects not only more complex and arguably precise diagnostic regimes, but has presaged the rise of more personalized treatments paradigms including targeted genomic therapies and immunotherapies for patients with tumors of the central nervous system. This shift in classification strategies and emerging cognate treatment paradigms represents an abrupt departure from historical approaches which relied categorically on histologic evaluation of specimens.
After providing this background and canvassing this historical context, I explore the ontological and epistemic upshots of these developments in neuro-oncology, which may be approached as a lense into the impact of genetics on the field of cancer biology as a whole. I explain that by furnishing a basis for imputing invariant features of neuro-oncologic reality, genomic criteria alter how cancer as a category is perceived and measured by key stakeholders, including patients, families, clinicians and resarchers. As the vocabulary used around cancer diagnoses has expanded to include genetic parameters, it has been argued that the identification and partitioning of diagnostic categories has become less vague, and increasingly numerous genuine classes, or natural types of cancer, have emerged. On the face of things, these next-generation methods (including chromosomal microarray analysis and next-generation sequencing) by which individuals come to know about the presense of absensce of oncologic entitites may be argued to have become less opaque and prone to more diagnostic certainty.
Despite their prima facie promise to clarify diagnostic regimes, however, I argue that emerging nosologic paradigms incorporating genomic features are complicated by increasing recognition that tumors of the central nervous system may be highly heterogeneous entities with potentially dissimilar genetic mutation patterns across cells within a given tumor. The inherently high degree of genomic instability within many such tumors that is conferred by disruption of cell-cycle regulatory mechanisms constitutive of cancer confounds reliable instantiation of uniform genomic properties. Given the current clinical standard wherein typically only a small part of a given tumor may be biopsied and gentically analyzed, intratumoral heteroneity, potential for clonal evolution and clonal expansion stand to obviate otherwise reliable epistemic approaches to uniformly classifying tumors by genetic properties or mutational status.
Rather than static, enduring properties, the genomic features of many tumors are prone to transform and transfigure. This is especially the case for tumors under aggressive treatment, for which longitudinal genomic and transcriptomic data has revealed high rates of clonal evolution wherein treatment ablates cells harboring susceptible genetic patterns and leaves behind cells with resistent genetic patterns. These findings have given rise to strategies to help differentiate driver mutations from carrier mutations.
As a way forward, a phylogenetic approach to conceptualizing and categorizing tumors, wherein intratumoral heterogeneity is explained by reference to heierarchical evolution away from a single clonal anscestor, can pave the way for a more coherent epistemic model for classifying cancer types, integrating genomic data without ignoring the inherent genomic instability harbored by these evolving entities.
Call for abstracts
The organizing committee welcomes abstracts for 20 minutes’ oral presentations on subjects that explore a problem with a conceptual, theoretical, methodological and/or philosophical approach and directly address questions relevant to cancer research.
Examples of questions that this workshop will raise include:
- How is the complexity of cancer addressed in scientific and medical practices?
- What are the epistemic implications of different approaches to cancer?
- How are questions about the ontology of cancer related to methodological issues?
- To what extent are questions concerning the nature of cancer intertwined with issues concerning diagnostics and/or treatments?
- How are new approaches, such as cancer genomics, organoids, and organ-on-chip technologies, influencing cancer research?
The call is open to philosophers, scientists and any person interested in the analysis of conceptual problems of cancer biology.
Abstract submission: Abstracts should be between 1000-1500 words long and must be submitted via this link before October 16th, 2019.
Notifications will be sent by the end of October.
Travel grants: PhD and postdoctoral scholars can apply for financial assistance to travel costs, up to 1000€ are awarded per person. In order to apply please submit your abstract via this link and send a letter of motivation to contact@philinbiomed.org.
Registration
The workshop is open to everyone, a small inscription fee of 50€ is asked for to help pay for the cost of organization and for food. Exemptions from the inscription fee can be granted upon demand, please write to contact@philinbiomed.com. In all cases registration is mandatory. Please sign up here: https://philocancer2.sciencesconf.org/registration
Organizing committee
Wiebke Bretting (ERC IDEM, ImmunoConcept, UMR5164, CNRS & University of Bordeaux)
Sara Green (Section for History and Philosophy of Science, Department of Science Education, University of Copenhagen, Denmark)
Lucie Laplane (IHPST, CNRS & University Paris 1; Institut Gustave Roussy, France)
Maël Lemoine (ImmunoConcept, UMR5164, CNRS & University of Bordeaux; IHPST, CNRS & University Paris 1, France)
Thomas Pradeu (ImmunoConcept, UMR5164, CNRS & University of Bordeaux; IHPST, CNRS & University Paris 1, France)
Scientific Committee
Sara Green (Section for History and Philosophy of Science, Department of Science Education, University of Copenhagen, Denmark)
Lucie Laplane (IHPST, CNRS & University Paris 1; Institut Gustave Roussy, France)
Maël Lemoine (ImmunoConcept, UMR5164, CNRS & University of Bordeaux, IHPST, CNRS & University Paris 1, France)
Jean-François Moreau (Professor Emeritus of Immunology, Univ. Bordeaux, Pellegrin Hospital & ImmunoConcept, Bordeaux, France).
Anya Plutynski (Department of Philosophy, Washington University in St. Louis, USA)
Thomas Pradeu (ImmunoConcept, UMR5164, CNRS & University of Bordeaux; IHPST, CNRS & University Paris 1, France)
Patronage Committee in Bordeaux
Andreas Bikfalvi (Professor of Biology and Director of the Angiogenesis and Cancer Microenvironment Laboratory, INSERM U1029, Univ. Bordeaux)
Bertrand Daignan-Fornier (Research Director in Genetics at CNRS, Head of IBGC, Bordeaux)
Julie Déchanet-Merville (Research Director in Immunology at CNRS, Head of ImmunoConcept, Bordeaux)
Jean-Luc Feugeas (Researcher in Applied Mathematics, Center for Intense Lasers and Applications, Bordeaux)
Guy Kantor (Professor of Medical Oncology and Radiotherapy, Institut Bergonié, Bordeaux)
Nicolas Larmonier (Professor of Immunology, Head of the research axis Cancer Immunology and Immunotherapies at ImmunoConcept, Bordeaux)
Simone Mathoulin-Pelissier (TBC) (Professor, Head of Team Epicene, Bordeaux Population Health)
Jean-François Moreau (Professor Emeritus of Immunology, Univ. Bordeaux, Pellegrin Hospital & ImmunoConcept, Bordeaux).
Violaine Moreau (Research Director at INSERM, BaRITON, Bordeaux Research in Translational Oncology, Bordeaux)
Maya Saleh (TBC) (Professor of Immunology at ImmunoConcept, Bordeaux)
Isabelle Sagot (Research Director at CNRS, IBGC, Bordeaux)
Frédéric Saltel (Research Director at INSERM, BaRITON, Bordeaux Research in Translational Oncology, Bordeaux)
David Santamaria (Group leader, Novel mediators in lung oncogenesis, IECB, Bordeaux)
Pierre Soubeyran (Head of research of Institut Bergonié, Comprehensive Cancer Center in Bordeaux, and Professor of Medical Oncology at the University of Bordeaux)
Martin Teichmann (Professor, Head of Transcription and Tumor Laboratory, University of Bordeaux)
Christine Varon (Professor, BaRITON, Bordeaux Research in Translational Oncology, Bordeaux)